Funding for Agri-food Data Canada is provided in part by the Canada First Research Excellence Fund
FAIR
Streamlining Data Documentation in Research
In of research, data documentation is often a complex and time-consuming task. To help researchers better document their data ADC has created the Semantic Engine as a powerful tool for creating structured, machine-readable data schemas. These schemas serve as blueprints that describe the various features and constraints of a dataset, making it easier to share, verify, and reuse data across projects and disciplines.
Defining Data
By guiding users through the process of defining their data in a standardized format, the Semantic Engine not only improves data clarity but also enhances interoperability and long-term usability. Researchers can specify the types of data they are working with, the descriptions of data elements, units of measurement used, and other rules that govern their values—all in a way that computers can easily interpret.
Introducing Range Overlays
With the next important update, the Semantic Engine now includes support for a new feature: range overlays.
Range overlays allow researchers to define expected value ranges for specific data fields, and if the values are inclusive or exclusive (e.g. up to but not including zero). This is particularly useful for quality control and verification. For example, if a dataset is expected to contain only positive values—such as measurements of temperature, population counts, or financial figures—the range overlay can be used to enforce this expectation. By specifying acceptable minimum and maximum values, researchers can quickly identify anomalies, catch data entry errors, and ensure their datasets meet predefined standards.
Verifying Data
In addition to enhancing schema definition, range overlay support has now been integrated into the Semantic Engine’s Data Verification tool. This means researchers can not only define expected value ranges in their schema, but also actively check their datasets against those ranges during the verification process.
When you upload your dataset into the Data Verification tool—everything running locally on your machine for privacy and security—you can quickly verify your data within your web browser. The tool scans each field for compliance with the defined range constraints and flags any values that fall outside the expected bounds. This makes it easy to identify and correct data quality issues early in the research workflow, without needing to write custom scripts or rely on external verification services.
Empowering Researchers to Ensure Data Quality
Whether you’re working with clinical measurements, survey responses, or experimental results, this feature lets you to catch outliers, prevent errors, and ensure your data adheres to the standards you’ve set—all in a user-friendly interface.
Written by Carly Huitema
Alrighty let’s briefly introduce this topic. AI or LLMs are the latest shiny object in the world of research and everyone wants to use it and create really cool things! I, myself, am just starting to drink the Kool-Aid by using CoPilot to clean up some of my writing – not these blog posts – obviously!!
Now, all these really cool AI tools or agents use data. You’ve all heard the saying “Garbage In…. Garbage Out…”? So, think about that for a moment. IF our students and researchers collect data and create little to no documentation with their data – then that data becomes available to an AI agent… how comfortable are you with the results? What are they based on? Data without documentation???
Let’s flip the conversation the other way now. Using AI agents for data creation or data analysis without understanding how the AI works, what it is using for its data, how do the models work – but throwing all those questions to the wind and using the AI agent results just the same. How do you think that will affect our research world?
I’m not going to dwell on these questions – but want to get them out there and have folks think about them. Agri-food Data Canada (ADC) has created data documentation tools that can easily fit into the AI world – let’s encourage everyone to document their data, build better data resources – that can then be used in developing AI agents.
![]()
At Agri-food Data Canada (ADC), we often emphasize the importance of content-derived identifiers—unique fingerprints generated from the actual content of a resource. These identifiers are especially valuable in research and data analysis, where reproducibility and long-term verification are essential. When you cite a resource using a derived identifier, such as a digest of source code, you’re ensuring that years down the line, anyone can confirm that the referenced material hasn’t changed.
One of the best tools for managing versioned documents—especially code—is GitHub. Not only does GitHub make it easy to track changes over time, but it also automatically generates derived identifiers every time you save your work.
What Is a GitHub Commit Digest?
Every time you make a commit in GitHub (i.e., save a snapshot of your code or document), GitHub creates a SHA-1 digest of that commit. This digest is a unique identifier derived from the content and metadata of the commit. It acts as a cryptographic fingerprint that ensures the integrity of the data.
Here’s what goes into creating a GitHub commit digest:
- Snapshot of the File Tree: Includes all file names, contents, and directory structure.
- Parent Commit(s): References to previous commits, which help maintain the history.
- Author Information: Name, email, and timestamp of the person who wrote the code.
- Committer Information: May differ from the author; includes who actually committed the change.
- Commit Message: The message describing the change.
All of this is bundled together and run through the SHA-1 hashing algorithm, producing a 40-character hexadecimal string like:
e68f7d3c9a4b8f1e2c3d4a5b6f7e8d9c0a1b2c3d
GitHub typically displays only the first 7–8 characters of this digest (e.g., e68f7d3c), which is usually enough to uniquely identify a commit within a repository.
Why You Should Reference Commit Digests
When citing code or documents stored on GitHub, always include the commit digest. This practice ensures that:
- Your references are precise and verifiable.
- Others can reproduce your work exactly as you did.
- The cited material remains unchanged and trustworthy, even years later.
Whether you’re publishing a paper, sharing an analysis, or collaborating on a project, referencing the commit digest helps maintain transparency and reproducibility, promoting FAIR research.
Final Thoughts
GitHub’s built-in support for derived identifiers makes it a powerful platform for version control and long-term citation. By simply noting the commit digest when referencing code or documents, you’re contributing to a more robust and verifiable research ecosystem.
So next time you cite GitHub work, take a moment to copy that digest. It’s a small step that makes a big difference in the integrity of your research.
Written by Carly Huitema
At Agri-food Data Canada (ADC), we are developing tools to help researchers create high-quality, machine-readable metadata. But what exactly is metadata, and what types does ADC work with?
What Is Metadata?
Metadata is essentially “data about data.” It provides context and meaning to data, making it easier to understand, interpret, and reuse. While the data itself doesn’t change, metadata describes its structure, content, and usage. Different organizations may define metadata slightly differently, depending on how they use it, but the core idea remains the same: metadata adds value by enhancing data context and improving the FAIRness of data.
Key Types of Metadata at ADC
At ADC, we focus on several types of metadata that are especially relevant to research outputs:
1. Catalogue Metadata
Catalogue metadata describes the general characteristics of a published work—such as the title, author(s), publication date, and publisher. If you’ve ever used a library card catalogue, you’ve interacted with this type of metadata. Similarly, when you cite a paper in your research, the citation includes catalogue metadata to help others locate the source.
2. Schema Metadata
Schema metadata provides detailed information about the structure and content of a dataset. It includes descriptions of variables, data formats, measurement units, and other relevant attributes. At ADC, we’ve developed a tool called the Semantic Engine to assist researchers in creating robust data schemas.
3. License Metadata
This type of metadata outlines the terms of use for a dataset, including permissions and restrictions. It ensures that users understand how the data can be legally accessed, shared, and reused.
These three types of metadata play a crucial role in supporting data discovery, interpretation, and responsible reuse.
Combining Metadata Types
Metadata types are not isolated—they often work together. For example, catalogue metadata typically follows a structured schema, such as Darwin Core, which itself has licensing terms (license metadata). Interestingly, Darwin Core is also catalogued: the Darwin Core schema specification has a title, authors, and a publication date.
– written by Carly Huitema
In our ongoing exploration of using the Semantic Engine to describe your data, there’s one concept we haven’t yet discussed—but it’s an important one: cardinality.
Cardinality refers to the number of values that a data field (specifically an array) can contain. It’s a way of describing how many items you’re expecting to appear in a given field, and it plays a crucial role in data descriptions, verification, and interpretation.
What Is an Array?
Before we talk about cardinality, we need to understand arrays. In data terms, an array is a field that can hold multiple values, rather than just one.
For example, imagine a dataset where you’re recording the languages a person speaks. Some people might speak only one language, while others might speak three or more. Instead of creating separate fields for “language1”, “language2”, and so on, you might store them all in one field as an array.
In an Excel spreadsheet, this might look like:
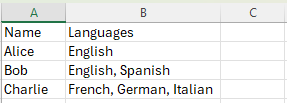
Written by Carly Huitema
I’ve been talking about data ownership in a few different ways and I have also been digging into these wonderful historical data sources. BUT! Where do we draw the line? Let’s be honest that’s another really tough question to answer.
I have proclaimed many times, to many different people/audiences, and in different fora – that I LOVE my historical data! I’ve asked the question to many – OAC is 151 years old now and we’ve been doing research for all those years – Where is all that data? Gone? Hidden? In someone’s basement?
Let me ask you this now… That historical data may be gone – but was there any value to it? Should we go out, find it, and make it accessible? Must ALL data be FAIR?? How does one decide?
I have NO clear answers to any of these questions – but I would love for you all to think about them. Is there truly any value to the data I collected during my BSc(Agr) degree? That was (Oh my! I’m going to say it) 39 years ago!!! Yes I still have the binder with all the rawdata – but should I do something with it? Should I make it FAIR?
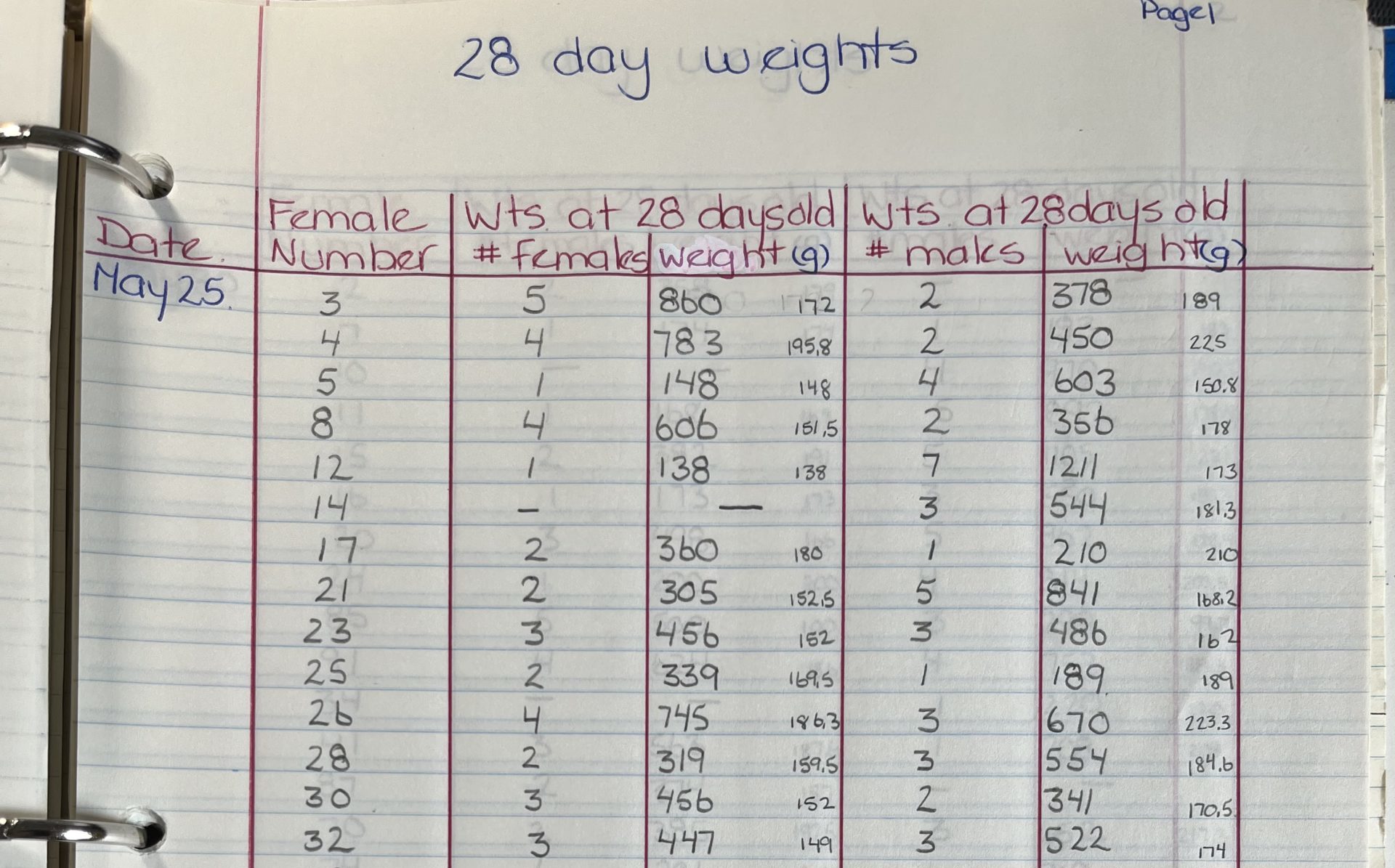
How do we determine if there is value? What do we keep and what do we throw away?
These are questions that archivists face everyday – but as a researcher – what do you think? Is the data you collected 10, 15, 20, 40 years ago have value? Should we make it FAIR???
I think you all know how I feel – if I had a magic wand – I would find and make all of our research data FAIR – doesn’t matter the age! For me, in the research context all data has value 🙂
![]()
In research and data-intensive environments, precision and clarity are critical. Yet one of the most common sources of confusion—often overlooked—is how units of measure are written and interpreted.
Take the unit micromolar, for example. Depending on the source, it might be written as uM, μM, umol/L, μmol/l, or umol-1. Each of these notations attempts to convey the same concentration unit. But when machines—or even humans—process large amounts of data across systems, this inconsistency introduces ambiguity and errors.
The role of standards
To ensure clarity, consistency, and interoperability, standardized units are essential. This is especially true in environments where data is:
-
Shared across labs or institutions
-
Processed by machines or algorithms
-
Reused or aggregated for meta-analysis
-
Integrated into digital infrastructures like knowledge graphs or semantic databases
Standardization ensures that “1 μM” in one dataset is understood exactly the same way in another and this ensures that data is FAIR (Findable, Accessible, Interoperable and Reusable).
UCUM: Unified Code for Units of Measure
One widely adopted system for encoding units is UCUM—the Unified Code for Units of Measure. Developed by the Regenstrief Institute, UCUM is designed to be unambiguous, machine-readable, compact, and internationally applicable.
In UCUM:
-
micromolar becomes
umol/L -
acre becomes
[acr_us] -
milligrams per deciliter becomes
mg/dL
This kind of clarity is vital when integrating data or automating analyses.
UCUM doesn’t include all units
While UCUM covers a broad range of units, it’s not exhaustive. Many disciplines use niche or domain-specific units that UCUM doesn’t yet describe. This can be a problem when strict adherence to UCUM would mean leaving out critical information or forcing awkward approximations. Furthermore, UCUM doesn’t offer and exhaustive list of all possible units, instead the UCUM specification describes rules for creating units. For the Semantic Engine we have adopted and extended existing lists of units to create a list of common units for agri-food which can be used by the Semantic Engine.
Unit framing overlays of the Semantic Engine
To bridge the gap between familiar, domain-specific unit expressions and standardized UCUM representations, the Semantic Engine supports what’s known as a unit framing overlay.
Here’s how it works:
-
Researchers can input units in a familiar format (e.g.,
acreoruM). -
Researchers can add a unit framing overlay which helps them map their units to UCUM codes (e.g.,
"[acr_us]"or"umol/L"). -
The result is data that is human-friendly, machine-readable, and standards-compliant—all at the same time.
This approach offers the both flexibility for researchers and consistency for machines.
Final thoughts
Standardized units aren’t just a technical detail—they’re a cornerstone of data reliability, semantic precision, and interoperability. Adopting standards like UCUM helps ensure that your data can be trusted, reused, and integrated with confidence.
By adopting unit framing overlays with UCUM, ADC enables data documentation that meet both the practical needs of researchers and the technical requirements of modern data infrastructure.
Written by Carly Huitema
Data schemas
Schemas are a type of metadata that provide context to your data, making it more FAIR (Findable, Accessible, Interoperable, and Reusable).
At their core, schemas describe your data, giving data better context. There are several ways to create a schema, ranging from simple to more complex. The simplest approach is to document what each column or field in your dataset represents. This can be done alongside your data, such as in a separate sheet within an Excel spreadsheet, or in a standalone text file, often referred to as a README or data dictionary.
However, schemas written as freeform text for human readers have limitations: they are not standardized and cannot be interpreted by machines. Machine-readable data descriptions offer significant advantages. Consider the difference between searching a library using paper card catalogs versus using a searchable digital database—machine-readable data descriptions bring similar improvements in efficiency and usability.
To enable machine-readability, various schema languages are available, including JSON Schema, JSON-LD, XML Schema, LinkML, Protobuf, RDF, and OCA. Each has unique strengths and use cases, but all allow users to describe their data in a standardized, machine-readable format. Once data is in such a format, it becomes much easier to convert between different schema types, enhancing its interoperability and utility.

Overlays Capture Architecture
The schema language Overlays Capture Architecture or OCA has two unique features which is why it is being used by the Semantic Engine.
- OCA embeds digests (specifically, OCA uses SAIDs)
- OCA is organized by features
Together these contribute to what makes OCA a unique and valuable way to document schemas.
OCA embeds digests
OCA uses digests which are digital fingerprints which can be used to unambiguously identify a schema. As digests are calculated directly from the content they identify this means that if you change the original content, the identifier (digest) also changes. Having a digital fingerprint calculated from the content is important for research reproducibility – it means you can find a digital object and if you have the identifier (digest) you can verify if the content has been changed. We have written a blog post about how digests are calculated and used in OCA.
OCA is organized by features
A schema describes the attributes of a dataset (typically the column headers) for a variety of features. A schema can be very simple and use very few features to describe attributes, but a more detailed schema will describe many features of each attribute.
We can represent a schema as a table, with rows for each attribute and a column for each feature.
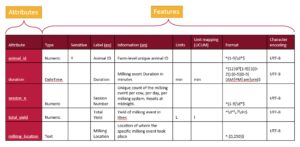
A tabular representation of a schema provides a clear overview of all the attributes and features used to describe a dataset. While schemas may be visualized as tables, they are ultimately saved and stored as text documents. The next step is to translate this tabular information into a structured text format that computers can understand.
From the table, there are two primary approaches to organizing the information in a text document. One method is to write it row by row, documenting an attribute followed by the values for each of its features. This approach, often called attribute-by-attribute documentation, is widely used in schema design such as JSON Schema and LinkML.

Schemas can also be written column by column, focusing on features instead of attributes. In this feature-by-feature approach (following the table in the figure above), you start by writing out all the data types for each attribute, then specify what is sensitive for each attribute, followed by providing labels, descriptions, and other metadata. These individual features, referred to as overlays in this schema architecture, offer a modular and flexible way to organize schema information. The Overlays Capture Architecture (OCA) is a global open overlay schema language that uses this method, enabling enhanced flexibility and modularity in schema design.

What is important for OCA is each overlay (feature) are given digests (the SAIDs). Each of the columns above is written out and a digest calculated and assigned, one digest for each feature. Then all the parts are put together and the entire schema is given a digest. In this way, all the content of a schema is bound together and it is never ambiguous about what is contained in a schema.
Why schema organization matters
Why does it matter whether schemas are written attribute-by-attribute or feature-by-feature? While we’ll explore this in greater detail in a future blog post, the distinction plays a critical role in calculating digests and managing governance in decentralized ecosystems.
A digest is a unique identifier for a piece of information, allowing it to be governed in a decentralized environment. When ecosystems of researchers and organizations agree on a specific digest (e.g., “version one schema” of an organization with digest xxx), they can agree on the schema’s validity and use.
A feature-by-feature schema architecture is particularly well-suited for governance. It offers flexibility by enabling individual features to be swapped, added, or edited without altering the core content of the data structure. Since the content remains unchanged, the digest also stays the same. This approach not only improves the schema’s adaptability but also enhances both the data’s and the schema’s FAIRness. This modularity ensures that schemas remain effective tools for collaboration and management in dynamic, decentralized ecosystems.
The Semantic Engine
All these details of an OCA schema are taken care of by the Semantic Engine. The Semantic Engine presents a user interface for generating a schema and writes the schema out in the language of OCA; feature by feature. The Semantic Engine calculates all the digests and puts them inside the schema document. It calculates the entire schema digest and it publishes that information when you export the schema. You can view all the digests (SAIDs) calculated for the schema in the readme.txt file.
Written by Carly Huitema
When you’re building a data schema you’re making decisions not only about what data to collect, but also how it should be structured. One of the most useful tools you have is format restrictions.
What Are Format Entries?
A format entry in a schema defines a specific pattern or structure that a piece of data must follow. For example:
- A date must look like YYYY-MM-DD or be in the ISO duration format.
- An email address must have the format name@example.com
- A DNA sequence might only include the letters A, T, G, and C
These formats are usually enforced using rules like regular expressions (regex) or standardized format types.
Why Would You Want to Restrict Format?
Restricting the format of data entries is about ensuring data quality, consistency, and usability. Here’s why it’s important:
✅ To Avoid Errors Early
If someone enters a date as “15/03/25” instead of “2025-03-15”, you might not know whether that’s March 15 or March 25 and what year? A clear format prevents confusion and catches errors before they become a problem.
✅ To Make Data Machine-Readable
Computers need consistency. A standardized format means data can be processed, compared, or validated automatically. For example, if every date follows the YYYY-MM-DD format, it’s easy to sort them chronologically or filter them by year. This is especially helpful for sorting files in folders on your computer.
✅ To Improve Interoperability
When data is shared across systems or platforms, shared formats ensure everyone understands it the same way. This is especially important in collaborative research.
Format in the Semantic Engine
Using the Semantic Engine you can add a format feature to your schema and describe what format you want the data to be entered in. While the schema writes the format rule in RegEx, you don’t need to learn how to do this. Instead, the Semantic Engine uses a set of prepared RegEx rules that users can select from. These are documented in the format GitHub repository where new format rules can be proposed by the community.
After you have created format rules in your schema you can use the Data Entry Web tool of the Semantic Engine to verify your results against your rules.
Final Thoughts
Format restrictions may seem technical, but they’re essential to building reliable, reusable, and clean data. When you use them thoughtfully, they help everyone—from data collectors to analysts—work more confidently and efficiently.
Written by Carly Huitema
In previous blog posts, we’ve discussed identifiers—specifically, derived identifiers, which are calculated directly from the digital content they represent. The key advantage of a derived identifier is that anyone can verify that the cited content is exactly what was intended. When you use a derived identifier, it ensures that the digital resource is authentic, no matter where it appears.
In contrast, authoritative identifiers work differently. They must be resolved through a trusted service, and you have to rely on that service to ensure the identifier hasn’t been altered and that the target hasn’t changed.
The Limitations of Derived Identifiers
One drawback of derived identifiers is that they only work for content that can be processed to generate a unique digest. Additionally, once an identifier is created, the content can no longer be updated. This can be a challenge when dealing with dynamic content, such as an evolving dataset or a standard that goes through multiple versions.
This brings us to the concept of identity, which goes beyond a simple identifier.
What Does Identity Mean?
Let’s take an example. The Global Alliance for Genomics and Health (GA4GH) publishes a data standard called Phenopackets. In this case Phenopackets is an identifier. Currently, there are two released versions (and two identifiers). However, anyone could create a new schema and call it “Phenopackets v3.” The key question is: is just naming something and giving it an identifier enough to have it be recognized as Phenopackets v3?
A name is not enough, what also matters is whether GA4GH itself releases “Phenopackets v3.” The identifier alone isn’t enough—we care about who endorses it. In this case, identity comes from GA4GH, the governing organization of Phenopackets.
Identity Through Reputation
Identity is established through reputation which is gained in two main ways:
- Transferred reputation – When an official organization (like GA4GH) endorses an identifier, the identity is backed by its authority and reputation.
- Acquired reputation – Even without a governing body, something gains identity via reputation if it becomes widely recognized and trusted.
For example, Bitcoin was created by an anonymous person (or group) using the pseudonym Satoshi Nakamoto—a name that doesn’t link to any legal identity which could grant it some reputation. Yet, the name Satoshi Nakamoto has strong identity via acquired reputation because of Bitcoin’s success and widespread recognition.
The key is that identity isn’t just about an identifier—it’s about who assigns it and why people trust it. To fully capture identity, we need to track not only the identifier but also the authority or reputation behind it.
How Do We Use an Identity?
Right now, we don’t have a universal system for identifying and verifying identity in a structured, machine-readable way. This is because identity is a combination of both an identifier and associated reputation/authority behind the identity and our current systems for identifiers don’t clearly recognize these two aspects of identity. Instead, we rely on indirect methods, like website URLs and domain names, to be a stand-in for the identity authority.
For example, if you want to verify the Phenopackets schema’s identity you would want to search out its associated authority. You might search for the Phenopackets name (the identifier) online or follow a link to its official GitHub repository. But how do you know that the GitHub page is legitimate? To confirm, you would check if the official GA4GH website links to it. Otherwise, anyone could create a GitHub repository and name it Phenopackets. The identifier is not enough, you also need to find the authority associated with the identity.
Another example of how we present the authority behind an identity are the academic journals. When they publish research, they add their reputation and peer-review process to build the reputation and identity of a paper. However, this system has flaws. When researchers cite papers they use DOIs which are specific identifiers of the journal article. The connection between the publication’s DOI to the identity of the paper is not standardized which makes discovery of important changes to the paper such as corrections and retractions challenging. Sometimes when you find the article on the journal webpage you might also find the retraction notice but this doesn’t always happen. This disconnect between identifiers and identity of publications leads to the proliferation of zombie publications which continue to be used even after they have been debunked.
Future Directions
As it stands, we lack effective tools for managing digital identity. This gap creates risks, including identity impersonation and difficulties in tracking updates, corrections, or retractions. Because our current citation system focuses on identifiers without strong linksing them to identity, important information can get lost. Efforts are underway to address these challenges, but we’re still in the early stages of finding solutions.
One early technology to address the growth of an identity has been Decentralized Identifiers (DIDs). We’ll talk more about them later, but they allow an identifier to be assigned to an identity that evolves and is provably under the control of an associated governance authority.
We hope this post has helped clarify the distinction between identifiers and identity which are often entangled — and why finding better ways to assign and verify identity is a problem worth solving.
Written by Carly Huitema



